150 years of TUM – Stories from the 2018 anniversary
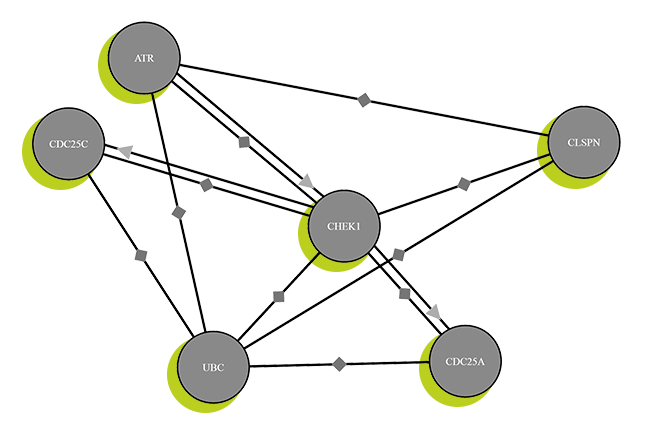
Protein map

The path to personalized medicine
Scrutinizing saliva, blood and earwax was all part of the quest to identify the multi-talented all-rounders of the human body: proteins. The mission also involved examining a wide variety of body tissues and fluids, and even numerous tumor cells. As a result of all this, in 2014, researchers led by biochemist Bernhard Küster were able to produce the first map of the human proteome – that is, the complete set of proteins inside our bodies. They successfully identified 18.097 of these proteins, which accounts for over 90 percent of the proteome. They also located these proteins within the body and worked out how many variants each one has.
This is an important step in developing personalized drug therapies, tailored specifically to each individual patient. In recent years it has become increasingly clear that not only our genes, but also our proteins have an influence on us and our diseases. This holds particularly true for cancer. Tumors develop from aberrant body cells, which vary greatly from one patient to another. Personalized medicine thus takes account of both the patient’s genetic makeup and their proteome. In the case of cancer, initial investigations by Küster and his research team suggest that the effect of drugs on cancer cells varies between people – precisely due to differences in proteins.
Filling in gaps around the globe
Küster and his team have used their results to create the most comprehensive protein database to date: ProteomicsDB. Newly developed software means they can store, link and evaluate their data – as can other protein experts all over the world.
Anyone can add the results of their own protein research, since the program is freely available on the Internet. In this way, all of the contributors are gradually filling in the last blanks on the human protein map.
Proteins – varied and versatile
Proteins are real masters of transformation, so analyzing them is extremely difficult. Depending on where they occur in the body and in what conditions, they slip into different roles, changing their chemical properties and appearance – which makes them even harder to differentiate.
So for each of the almost 20,000 proteins identified by Küster and his team, there are still countless variants. The task facing researchers in the future is to find them all.
Key tool: the mass spectrometer
Since proteins are tiny, advanced analytical techniques are required to study them. One such technique is mass spectrometry. Here, a protein’s molecular chain is first broken down into short fragments, which are sorted according to weight and electrical charge. The data obtained from these individual pieces can then be evaluated and virtually reassembled by computer – similar to doing a jigsaw puzzle.
By comparing this data with established data sets, researchers can then determine very precisely which protein is involved. This process thus depends on large databases containing all known proteins – such as the ProteomicsDB set up by Bernhard Küster.
“If we can establish the protein profile of a tumor, for instance, we could administer drugs in a much more selective way in the future.”
Bernhard Küster, Professor of Proteomics and Bioanalytics at TUM
Disclaimer
This story was published in 2018 to mark TUM’s 150th anniversary on a jubilee website that has since been deactivated.
Text: TUM Web Communications Team; Graphics: KW NEUN
Literature on the history of TUM
- Wolfgang A. Herrmann (Hrsg.), Martin Pabst/Margot Fuchs (Verf.), Technische Universität München - Geschichte eines Wissenschaftsunternehmens, 2 Bd., Berlin 2006.
Link to the online catalog of the University Library - Wolfgang A. Herrmann, Winfried Nerdinger (Hrsg.), Die Technische Hochschule München im Nationalsozialismus, München 2018.
Link to free download via mediaTUM (PDF, in German, 79 MB)
Link to order the book
Link to copies in the University Library - Irene Meissner, Bauten+Kunst. Technische Universität München 1868-2018, München 2018. Link to the online catalog of the University Library
- Martin Pabst, Alumni der TUM. Prägende Gestalter aus der Technischen Universität München, München 2018. Link to the online catalog of the University Library
- Martin Pabst, Köpfe der TUM. Geniale Entdecker und Erfinder aus der Technischen Universität München, München 2018. Link to the online catalog of the University Library
- Brigitte Röthlein, Pioniere gestalten die Welt der Technik. 150 Jahre Forschung an der Technischen Universität München, München 2018. Link to the online catalog of the University Library
Further books and information on the history of TUM
Acknowledgements
We would like to thank everyone who helped us write the texts and create the visualizations. In particular, we would like to thank the authors of the books mentioned, the experts at the chairs, professors, staff, and press officers at the TUM Corporate Communications Center. We would also like to thank the staff of the Architecture Museum, the TUM German Heart Center, the TUM Klinikum rechts der Isar, the European Space Agency (ESA), and everyone else who provided us with expert advice and image material.
The anniversary stories were written by the TUM Web Communications team. The graphic content was created by KW NEUN – Designagentur.